End-to-end examples (MToolBox)
Prerequisites Installation, reference bundles, and all dependencies must be completed beforehand.
Architecture MToolBox supports x86_64 only. ARM-based systems, including Apple Silicon (M1/M2/M3), are not supported.
- MIT single-sample run
- MIT cohort run
1. Before running mtDNA calling you must have a BAM file from WES/WGS
Does it matter if I ran WES/WGS with GATK 3.5 or GATK 4.6? No. CBIcall will detect and use the
bamfiles produced by either version.
Just make sure thatbamfiles are available — FASTQ input is not supported.
CBIcall expects a BAM file from a previous WES/WGS run:
CNAG999_exome
└── CNAG99901P_ex <--- ID taken from here
└── *cbicall_bash_w?s_single_gatk-* <- The script expects that you have a BAM file inside this directory
Note on nomenclature Please see this page.
2. Create a parameters file
Create a YAML file, e.g. mit_single.yaml.
Important Please make sure you use the same value for the key
samplethat you used for WES/WGS.
Example:
mode: single
pipeline: mit
workflow_engine: bash
input_dir: CNAG999_exome/CNAG99901P_ex
See Configuration Reference for all YAML keys and supported combinations.
3. Run CBIcall
bin/cbicall run -p mit_single.yaml -t 4
-pselects the YAML parameters file-tsets the number of threads
4. Inspect outputs
After completion, you will find:
CNAG999_exome/CNAG99901P_ex/cbicall_bash_mit_single_rsrs_gatk-3.5_*/
01_mtoolbox/
02_browser/
- Final mtDNA report:
01_mtoolbox/mit_prioritized_variants.txt - mtDNA VCF:
01_mtoolbox/VCF_file.vcf - Browser report:
02_browser/<run-id>.html
See Outputs for the full file reference.
5. Visualize variants in the browser
Please see:
02_browser/README.txt
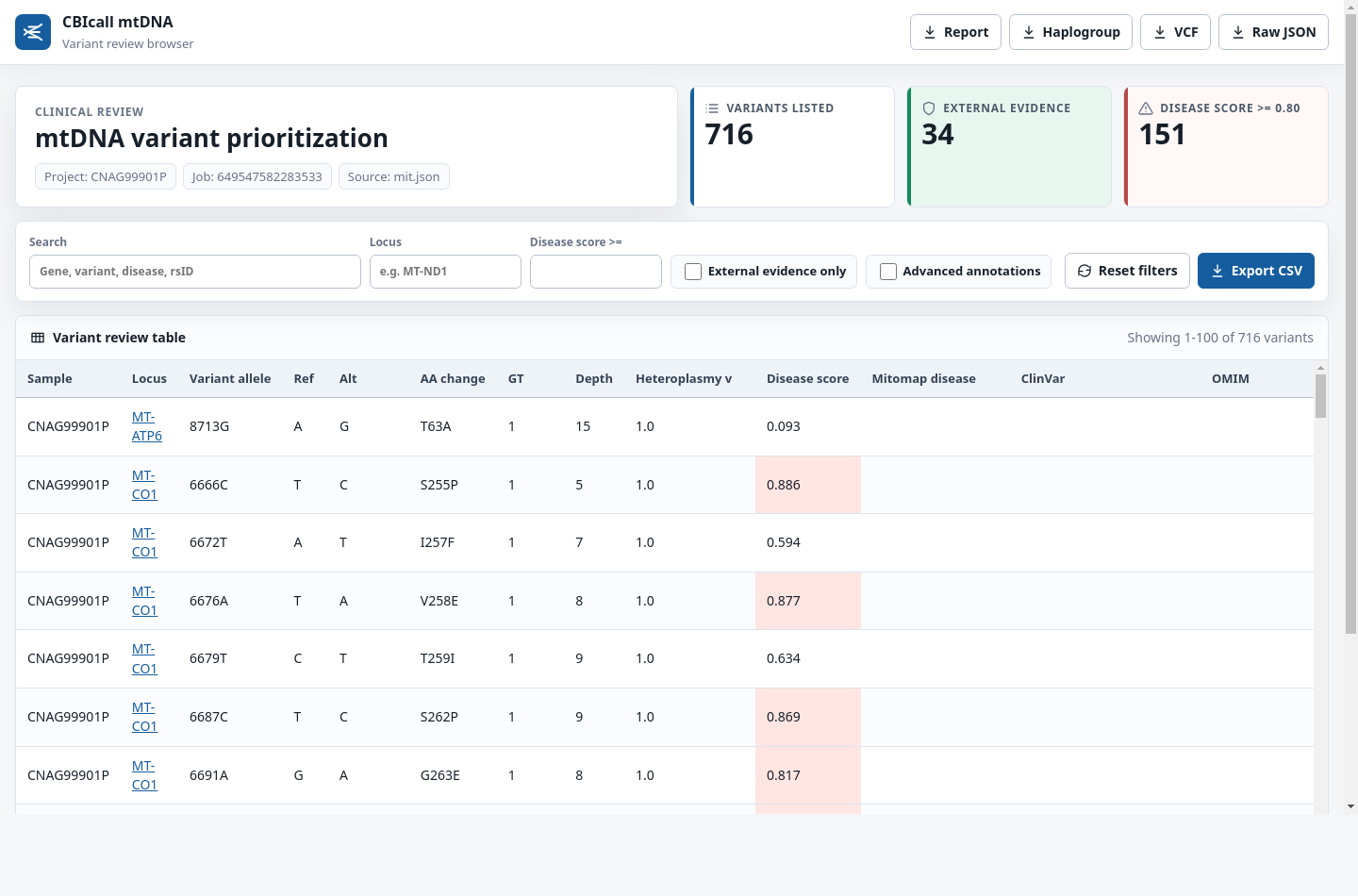
The CBIcall mtDNA variation browser is a standalone HTML report. It embeds the
browser payload at generation time, so 02_browser/<run-id>.html can be opened
directly without a local web server or external static assets.
See snapshot
Browser actions
The report provides direct buttons for:
- Report:
01_mtoolbox/mit_prioritized_variants.txt, including annotations plus appendedGT,DP, and heteroplasmy values. - Haplogroup:
01_mtoolbox/mt_classification_best_results.csv, including the predicted haplogroup for each sample. - VCF:
01_mtoolbox/VCF_file.vcf, containing the mtDNA variants in VCF format. - Raw JSON:
01_mtoolbox/mit.raw.json, containing the unfiltered parsed MToolBox output.
The browser also supports searching by gene, variant, disease term, or rsID; filtering by locus; filtering by minimum disease score; showing only variants with external evidence; toggling advanced annotation columns; and exporting the current table view as CSV.
HTML table:
The CBIcall mtDNA variation browser displays a browsable table consisting of the most relevant fields relative to the variant annotation:
- Sample: The full name of each sample.
- Locus: The location on the mitochondrial chromosome.
- Variant allele: The position in the mitochondrial chromosome + the alternative allele format.
- Ref: The reference allele (mitochondrial reference genome: RSRS).
- Alt: The alternative allele(s).
- AA change: The amino acid change if the variant falls in a coding region.
- GT: Genotype. 0:Ref, ≥1:Alt(s).
- Depth: The number of times this position is covered by reads.
- Heteroplasmy: The heteroplasmic fraction. Note that the confidence interval can be retrieved from the downloadable VCF file.
- Other: For other fields please consult MToolBox's manual.
Filtered variants The table shows pre-filtered variants. Variants were excluded if:
- HF ≤ 0.30 (maximum HF observed in any sample)
- 1000 Genomes frequency ≥ 0.01
- Not present in the input VCF
By default, variants with missing HF values (NA,N/A,.) are excluded.
Use the --keep-missing-hf option to retain them.
For advanced parameters, multi-sample analyses, mtDNA workflows and troubleshooting, see the Usage and FAQ sections.
1. Before running mtDNA calling you must have BAM files from WES/WGS
Does it matter if I ran WES/WGS with GATK 3.5 or GATK 4.6? No. CBIcall will detect and use the
bamfiles produced by either version.
Just make sure thatbamfiles are available — FASTQ input is not supported.
CBIcall expects BAM files from previous WES/WGS runs:
CNAG999_exome
└── CNAG99901P_ex <--- ID taken from here
└── *cbicall_bash_w?s_single_gatk-* <- The script expects that you have a BAM file inside this directory
CNAG99902M_ex <--- ID taken from here
└── *cbicall_bash_w?s_single_gatk-* <- The script expects that you have a BAM file inside this directory
Note on nomenclature Please see this page.
2. Create a parameters file
Create a YAML file, e.g. mit_cohort.yaml:
mode: cohort
pipeline: mit
workflow_engine: bash
gatk_version: gatk-3.5
input_dir: CNAG999_exome
See Configuration Reference for all YAML keys and supported combinations.
3. Run CBIcall
bin/cbicall run -p mit_cohort.yaml -t 4
-pselects the YAML parameters file-tsets the number of threads
4. Inspect outputs
After completion, you will find:
CNAG999_exome/cbicall_bash_mit_cohort_rsrs_gatk-3.5*
01_mtoolbox/
02_browser/
- Final joint mtDNA report:
01_mtoolbox/mit_prioritized_variants.txt - Joint mtDNA VCF:
01_mtoolbox/VCF_file.vcf - Browser report:
02_browser/<run-id>.html
See Outputs for the full file reference.
5. Visualize variants in the browser
Please see:
02_browser/README.txt
The CBIcall mtDNA variation browser is a standalone HTML report. It embeds the
browser payload at generation time, so 02_browser/<run-id>.html can be opened
directly without a local web server or external static assets. The cohort report
follows the same standalone HTML format as the single-sample report, with
sample-level fields where applicable.
Browser actions
The report provides direct buttons for:
- Report:
01_mtoolbox/mit_prioritized_variants.txt, including annotations plus appended per-sampleGT,DP, and heteroplasmy values. - Haplogroup:
01_mtoolbox/mt_classification_best_results.csv, including the predicted haplogroup for each sample. - VCF:
01_mtoolbox/VCF_file.vcf, containing the mtDNA variants in VCF format. - Raw JSON:
01_mtoolbox/mit.raw.json, containing the unfiltered parsed MToolBox output.
The browser also supports searching by gene, variant, disease term, or rsID; filtering by locus; filtering by minimum disease score; showing only variants with external evidence; toggling advanced annotation columns; and exporting the current table view as CSV.
HTML table:
The CBIcall mtDNA variation browser displays a browsable table consisting of the most relevant fields relative to the variant annotation:
- Sample: The full name of each sample.
- Locus: The location on the mitochondrial chromosome.
- Variant allele: The position in the mitochondrial chromosome + the alternative allele format.
- Ref: The reference allele (mitochondrial reference genome: RSRS).
- Alt: The alternative allele(s).
- AA change: The amino acid change if the variant falls in a coding region.
- GT: Genotype. 0:Ref, ≥1:Alt(s).
- Depth: The number of times this position is covered by reads.
- Heteroplasmy: The heteroplasmic fraction. Note that the confidence interval can be retrieved from the downloadable VCF file.
- Other: For other fields please consult MToolBox's manual.
Filtered variants The table shows pre-filtered variants. Variants were excluded if:
- HF ≤ 0.30 (maximum HF observed in any sample)
- 1000 Genomes frequency ≥ 0.01
- Not present in the input VCF
By default, variants with missing HF values (NA,N/A,.) are excluded.
Use the --keep-missing-hf option to retain them.
Genetic data interpretation disclaimer: review the project disclaimer before clinical or diagnostic interpretation.