Architecture
CBIcall is a thin orchestration layer around one or more concrete pipelines. Its main responsibilities are:
- Reading a parameters YAML file
- Validating required parameters and paths
- Resolving the selected pipeline and workflow engine
- Preparing the project directory structure
- Calling the appropriate workflow scripts (Bash or Snakemake)
- Managing logs and collecting results in a standard layout
The actual bioinformatics work (alignment, variant calling, QC) is implemented in modular pipelines that can be extended or replaced.
Architecture diagram
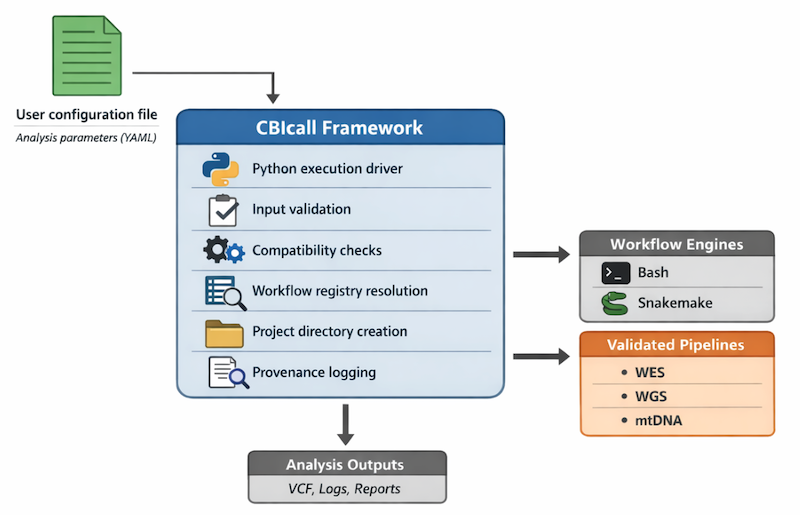
CBIcall resolves a validated parameters YAML into a concrete workflow engine and pipeline implementation.
Main components
At a high level, CBIcall consists of:
Execution driver
Reads YAML, validates parameters, builds the run directory, and dispatches the selected workflow.
Workflow resolver
Maps engine, GATK version, pipeline, and mode to a concrete script or Snakefile.
Analysis layer
Runs alignment, variant calling, mtDNA analysis, QC, and report generation.
Outputs
Stores BAMs, VCFs, stats, browser reports, logs, and the resolved run metadata.
| Component | Role | Main files or directories |
|---|---|---|
| Python execution driver | Reads the YAML configuration, validates parameters, resolves paths, and dispatches execution to the selected workflow. | src/cbicall/config.py, src/cbicall/dnaseq.py |
| Workflow registry | Developer-facing map that connects parameters YAML choices (workflow_engine, gatk_version, pipeline, mode, and pipeline implementation version) to concrete workflow scripts. Validate it with bin/cbicall validate-registry after editing. | workflows/registry/workflows.yaml, src/cbicall/workflow_registry.py |
| Pipelines | Implement WES, WGS, and mtDNA analyses. A pipeline may provide Bash workflows, Snakemake workflows, or both. | workflows/bash/, workflows/snakemake/ |
| Workflow engines | Execute the resolved workflow. Bash is transparent and direct; Snakemake adds rule-based orchestration and partial targets. | BashRunner, SnakemakeRunner in src/cbicall/dnaseq.py |
| Run directory | Stores outputs, logs, and log.json for one execution. | 01_bam/, 02_varcall/, 03_stats/, logs/ |
| External data | Provides third-party tools, reference genomes, known-sites resources, and accessory databases. | DATADIR, NGSUTILS, Databases |
Directory structure
<project_dir>/
01_bam/
02_varcall/
03_stats/
logs/
Typical usage:
- Intermediate alignment and BAM files are stored under
01_bam/. - Variant-calling outputs (gVCFs, VCFs and related files) are stored under
02_varcall/. - Summary statistics and QC metrics are collected under
03_stats/. - Log files for all steps are stored under
logs/.
Execution model
CBIcall supports two execution modes:
-
Single mode
Each sample is processed independently. -
Cohort mode
Joint analysis using per-sample gVCFs from previous single runs.
The workflow engine is selected in the YAML:
workflow_engine: bashworkflow_engine: snakemake
Rule of thumb: Bash workflows are direct and transparent. Snakemake workflows are better when rule-based orchestration or partial targets matter.
Supported pipelines
| Pipeline | Mode | Genome | GATK version | Status / Notes |
|---|---|---|---|---|
| WES | single | b37 (default) | gatk-3.5, gatk-4.6 | ✓ Supported |
| WES | cohort | b37 (default) | gatk-3.5, gatk-4.6 | ✓ Supported |
| WGS | single | b37 (default), hg38 | gatk-4.6 | ✓ Supported |
| WGS | cohort | b37 (default), hg38 | gatk-4.6 | ✓ Supported |
| MIT (mtDNA) | single | rsrs (fixed) | gatk-3.5 | ⚠ Not supported on ARM / aarch64 |
| MIT (mtDNA) | cohort | rsrs (fixed) | gatk-3.5 | ⚠ Not supported on ARM / aarch64 |
✓ Fully supported configuration ⚠ Platform limitation
Date: March-2026
Extensibility
New pipelines can be added without modifying the core system:
- Each pipeline lives under
workflows/<engine>/<name> - The execution driver resolves the pipeline implementation through the workflow registry
- Pipelines reuse the same directory layout and logging conventions
- Pipelines may support single and/or cohort mode
See: