What is Convert-Pheno?¶
Convert-Pheno is an open-source toolkit for converting clinical and phenotypic data between several commonly used exchange models.
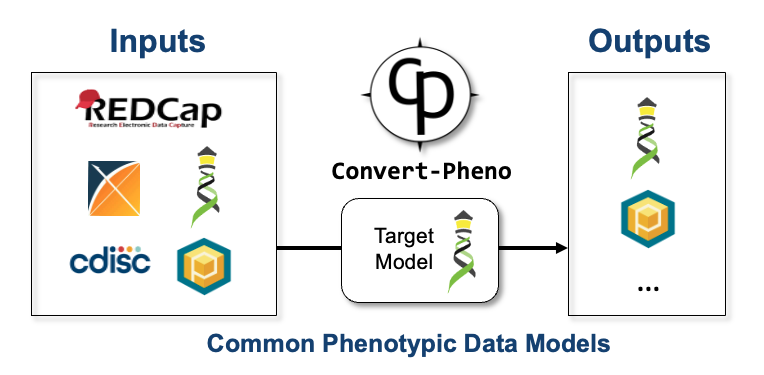
In practice, the project is centered on a Perl module and a command-line tool that work with text files such as:
- BFF
individualsJSON/YAML, plus entity-aware BFF output forbiosamples,datasets, andcohorts - Phenopackets v2 JSON/YAML
- OMOP-CDM SQL or CSV exports
- experimental openEHR canonical JSON/YAML composition input
- REDCap CSV exports
- CDISC-ODM XML
- mapped CSV input
Internally, the toolkit uses BFF as its center model for most conversions. From there, it can emit BFF, PXF, or OMOP CDM output depending on the selected workflow.
The current openEHR path should still be treated as experimental. It is currently aimed at canonical composition input and BFF or PXF output.
Most users should start here¶
For most users, the command-line interface is the right entry point. It is the most direct way to run conversions on files and the interface that the rest of the project is built around.
Typical examples are:
convert-pheno -ipxf pxf.json -obff individuals.json
convert-pheno -ipxf pxf.json -obff --entities individuals biosamples datasets cohorts --out-dir out/
convert-pheno -iomop dump.sql.gz -obff individuals.json.gz --stream --ohdsi-db
See Use as a command-line interface for the CLI entry points and Usage for more examples.
Other ways to use it¶
If you need tighter integration:
- As a module: call
Convert::Phenofrom Perl code - As an API: run the lightweight HTTP API
- Web App UI: inspect and try conversions interactively
What to read next¶
- Supported formats if you want to know what can be converted
- Download & Installation if you want to install it
- Use as a command-line interface if you want to run it now