Beacon v2 Models (BFF)
BFF stands for Beacon Friendly Format. The BFF is a data exchange format composed of 7 JSON files. These files correspond to the 7 entities of the Beacon v2 Models.
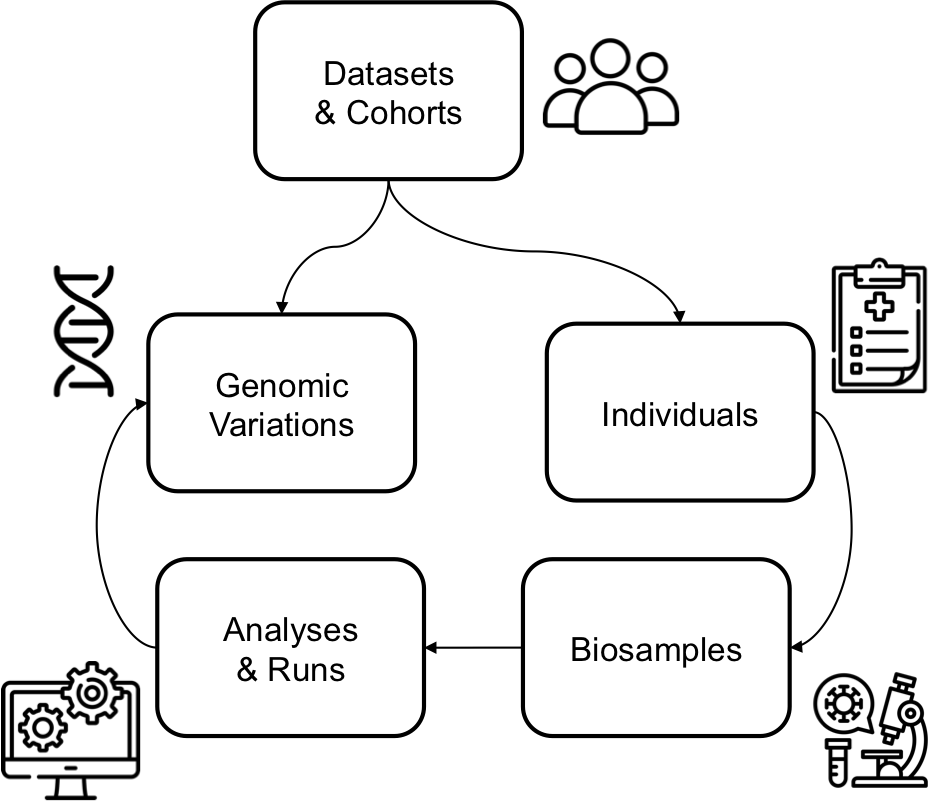
About Beacon v2 Models' entities
Among the Beacon v2 Models entities, individuals is usually the main record-level carrier of phenotypic and clinical information. Other entities such as biosamples, datasets, and cohorts are also important, but they tend to be more focused: biosamples capture specimen-level context, while datasets and cohorts are mostly collection-level metadata. In practice, this means individuals usually drives the most complex semantic mapping, whereas the other entities are often derived, enriched, or populated with lighter transformation rules.
Convert-Pheno supports BFF in both directions.
- As input, it currently accepts the individuals entity serialized as
individuals.json. - As output, the CLI can emit
individuals,biosamples,datasets, andcohorts.
So BFF is no longer only an individuals-centered output format, even though reverse conversion from BFF input is still centered on individuals.
Browsing BFF JSON data
You can browse a public BFF v2 file with the following JSON viewers:
BFF As Input  ¶
¶
When using the convert-pheno command-line interface, simply ensure the correct syntax is provided.
About JSON data in individuals.json
If the file individuals.json is a JSON array of objects (for which each object corresponds to an individual), the output -opxf file will also be a JSON array.
The module interface takes one flat payload. Unlike the HTTP API, module arguments are not split into input, output, and options.
Send a POST request to the API URL (see more info here) with a small payload like:
{
"conversion": "bff2pxf",
"input": {
"data": {
"id": "HG00096",
"sex": {
"id": "NCIT:C20197",
"label": "male"
}
}
},
"output": {
"entities": ["individuals"]
}
}
Successful API responses wrap the conversion result under data.
BFF As Output¶
As an output format, BFF can be emitted in two modes:
- individuals-only BFF mode, using
-obff FILE - entity-aware mode, using
--entities ... --out-dir DIR, which can write multiple Beacon entities
The currently supported BFF output entities are:
individualsbiosamplesdatasetscohorts
Examples:
convert-pheno -ipxf pxf.json -obff individuals.json
convert-pheno -ipxf pxf.json --entities individuals biosamples --out-dir out/
convert-pheno -icsv clinical.csv --mapping-file clinical.yaml --entities individuals datasets cohorts --out-dir out/
At the moment, reverse conversion from BFF input still expects individuals rather than a full multi-entity Beacon bundle.
Please find below examples of data:
{
"ethnicity": {
"id": "NCIT:C42331",
"label": "African"
},
"id": "HG00096",
"info": {
"eid": "fake1"
},
"interventionsOrProcedures": [
{
"procedureCode": {
"id": "OPCS4:L46.3",
"label": "OPCS(v4-0.0):Ligation of visceral branch of abdominal aorta NEC"
}
}
],
"measures": [
{
"assayCode": {
"id": "LOINC:35925-4",
"label": "BMI"
},
"date": "2021-09-24",
"measurementValue": {
"quantity": {
"unit": {
"id": "NCIT:C49671",
"label": "Kilogram per Square Meter"
},
"value": 26.63838307
}
}
},
{
"assayCode": {
"id": "LOINC:3141-9",
"label": "Weight"
},
"date": "2021-09-24",
"measurementValue": {
"quantity": {
"unit": {
"id": "NCIT:C28252",
"label": "Kilogram"
},
"value": 85.6358
}
}
},
{
"assayCode": {
"id": "LOINC:8308-9",
"label": "Height-standing"
},
"date": "2021-09-24",
"measurementValue": {
"quantity": {
"unit": {
"id": "NCIT:C49668",
"label": "Centimeter"
},
"value": 179.2973
}
}
}
],
"sex": {
"id": "NCIT:C20197",
"label": "male"
}
}
{
"diseases" : [],
"id" : "phenopacket_id.AUNb6vNX1",
"measurements" : [
{
"assay" : {
"id" : "LOINC:35925-4",
"label" : "BMI"
},
"value" : {
"quantity" : {
"unit" : {
"id" : "NCIT:C49671",
"label" : "Kilogram per Square Meter"
},
"value" : 26.63838307
}
}
},
{
"assay" : {
"id" : "LOINC:3141-9",
"label" : "Weight"
},
"value" : {
"quantity" : {
"unit" : {
"id" : "NCIT:C28252",
"label" : "Kilogram"
},
"value" : 85.6358
}
}
},
{
"assay" : {
"id" : "LOINC:8308-9",
"label" : "Height-standing"
},
"value" : {
"quantity" : {
"unit" : {
"id" : "NCIT:C49668",
"label" : "Centimeter"
},
"value" : 179.2973
}
}
}
],
"medicalActions" : [
{
"procedure" : {
"code" : {
"id" : "OPCS4:L46.3",
"label" : "OPCS(v4-0.0):Ligation of visceral branch of abdominal aorta NEC"
},
"performed" : {
"timestamp" : "1900-01-01T00:00:00Z"
}
}
}
],
"metaData" : null,
"subject" : {
"id" : "HG00096",
"sex" : "MALE",
"vitalStatus" : {
"status" : "ALIVE"
}
}
}