Cohort Mode
Cohort mode performs an all-vs-all comparison of records in one or more cohorts. Each record is flattened, encoded as a binary vector, and compared with either Hamming distance or the Jaccard index.
Use cohort mode when you want to explore the structure of a cohort, compare multiple cohorts, identify clusters, run dimensionality reduction, or export a graph for network analysis.
pheno-ranker -r cohort.jsonmatrix.txtWhen to Use It
Explore
Cohort structure
Generate pairwise distances or similarities between all records in a cohort.
Compare
Multiple cohorts
Pass several reference files and keep source cohorts traceable with prefixes.
Scale
Large outputs
Use sparse Matrix Market output or graph edge filters when dense outputs are too large.
Inspect
Intermediate vectors
Export hashes, binary vectors, and coverage statistics for debugging or downstream use.
What You Get
matrix.txt: the default dense pairwise comparison matrix.graph.json: an optional Cytoscape-compatible graph when--cytoscape-jsonis used.graph_stats.txt: optional graph summary statistics when--graph-statsis used.export.*.json: optional intermediate hashes, vectors, and coverage statistics when--exportis used.matrix.mtx: optional sparse Matrix Market output for large matrix workflows.
Use cohort mode when every record should be compared with every other record. Use patient mode when you have a target patient or object and want a ranked list of closest matches.
We created a tutorial that deliberately uses generic JSON data (i.e., movies) to illustrate the capabilities of Pheno-Ranker, as starting with familiar examples can help you better grasp its usage.
Once you are comfortable with the concepts using movie data, you will find it easier to apply Pheno-Ranker to real GA4GH standards. For specific examples, please refer to the cohort and patient pages in this documentation.
Usage
The examples below show common cohort-mode command-line patterns. For the complete CLI reference, see Usage.
- Intra-cohort
- Inter-cohort
For this example, we use individuals.json, a JSON array with 36 patients. The goal is to compare every patient against every other patient in the file.
First, we will download the file:
wget https://raw.githubusercontent.com/CNAG-Biomedical-Informatics/pheno-ranker/refs/heads/main/t/data/individuals.json
Now run Pheno-Ranker:
pheno-ranker -r individuals.json
More input examples
You can find more input examples here.
This process generates a matrix.txt file, containing the results of 36 x 36 pairwise comparisons, calculated using the Hamming distance metric.
See matrix.txt
| 107:week_0_arm_1 | 107:week_2_arm_1 | 107:week_14_arm_1 | 125:week_0_arm_1 | 125:week_2_arm_1 | 125:week_14_arm_1 | 125:week_26_arm_1 | 125:week_52_arm_1 | 125:week_78_arm_1 | 215:week_0_arm_1 | 215:week_2_arm_1 | 215:week_14_arm_1 | 215:week_26_arm_1 | 215:week_52_arm_1 | 215:week_78_arm_1 | 257:week_0_arm_1 | 257:week_2_arm_1 | 257:week_14_arm_1 | 257:week_26_arm_1 | 275:week_0_arm_1 | 275:week_2_arm_1 | 275:week_14_arm_1 | 275:week_52_arm_1 | 305:week_0_arm_1 | 305:week_26_arm_1 | 305:week_52_arm_1 | 365:week_0_arm_1 | 365:week_2_arm_1 | 365:week_14_arm_1 | 365:week_26_arm_1 | 365:week_52_arm_1 | 527:week_0_arm_1 | 527:week_2_arm_1 | 527:week_14_arm_1 | 527:week_26_arm_1 | 527:week_52_arm_1 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 107:week_0_arm_1 | 0 | 24 | 23 | 6 | 23 | 23 | 24 | 43 | 40 | 16 | 27 | 29 | 27 | 49 | 32 | 29 | 45 | 45 | 50 | 14 | 25 | 30 | 51 | 18 | 26 | 45 | 20 | 25 | 26 | 30 | 45 | 24 | 24 | 23 | 32 | 43 |
| 107:week_2_arm_1 | 24 | 0 | 3 | 22 | 3 | 3 | 2 | 23 | 18 | 30 | 7 | 9 | 7 | 29 | 10 | 47 | 25 | 25 | 28 | 30 | 5 | 10 | 31 | 32 | 4 | 25 | 34 | 5 | 6 | 8 | 25 | 42 | 2 | 3 | 10 | 23 |
| 107:week_14_arm_1 | 23 | 3 | 0 | 21 | 2 | 2 | 3 | 22 | 19 | 29 | 6 | 8 | 6 | 28 | 11 | 46 | 24 | 24 | 29 | 29 | 4 | 9 | 30 | 31 | 5 | 24 | 33 | 4 | 5 | 9 | 24 | 41 | 3 | 2 | 11 | 22 |
| 125:week_0_arm_1 | 6 | 22 | 21 | 0 | 21 | 21 | 22 | 41 | 38 | 14 | 25 | 27 | 25 | 47 | 30 | 29 | 43 | 43 | 48 | 12 | 23 | 28 | 49 | 16 | 24 | 43 | 18 | 23 | 24 | 28 | 43 | 24 | 22 | 21 | 30 | 41 |
| 125:week_2_arm_1 | 23 | 3 | 2 | 21 | 0 | 2 | 3 | 22 | 19 | 29 | 6 | 8 | 6 | 28 | 11 | 46 | 24 | 24 | 29 | 29 | 4 | 9 | 30 | 31 | 5 | 24 | 33 | 4 | 5 | 9 | 24 | 41 | 3 | 2 | 11 | 22 |
| 125:week_14_arm_1 | 23 | 3 | 2 | 21 | 2 | 0 | 3 | 22 | 19 | 29 | 6 | 8 | 6 | 28 | 11 | 46 | 24 | 24 | 29 | 29 | 4 | 9 | 30 | 31 | 5 | 24 | 33 | 4 | 5 | 9 | 24 | 41 | 3 | 2 | 11 | 22 |
| 125:week_26_arm_1 | 24 | 2 | 3 | 22 | 3 | 3 | 0 | 23 | 18 | 30 | 7 | 9 | 7 | 29 | 10 | 47 | 25 | 25 | 28 | 30 | 5 | 10 | 31 | 32 | 4 | 25 | 34 | 5 | 6 | 8 | 25 | 42 | 2 | 3 | 10 | 23 |
| 125:week_52_arm_1 | 43 | 23 | 22 | 41 | 22 | 22 | 23 | 0 | 7 | 49 | 26 | 28 | 26 | 8 | 15 | 26 | 4 | 4 | 9 | 49 | 24 | 29 | 10 | 51 | 25 | 4 | 53 | 24 | 25 | 29 | 4 | 21 | 23 | 22 | 15 | 2 |
| 125:week_78_arm_1 | 40 | 18 | 19 | 38 | 19 | 19 | 18 | 7 | 0 | 46 | 23 | 25 | 23 | 13 | 10 | 31 | 9 | 9 | 12 | 46 | 21 | 26 | 15 | 48 | 20 | 9 | 50 | 21 | 22 | 24 | 9 | 26 | 18 | 19 | 10 | 7 |
| 215:week_0_arm_1 | 16 | 30 | 29 | 14 | 29 | 29 | 30 | 49 | 46 | 0 | 33 | 27 | 33 | 43 | 38 | 37 | 51 | 51 | 56 | 12 | 31 | 34 | 45 | 22 | 32 | 51 | 18 | 31 | 30 | 36 | 51 | 34 | 30 | 29 | 38 | 49 |
| 215:week_2_arm_1 | 27 | 7 | 6 | 25 | 6 | 6 | 7 | 26 | 23 | 33 | 0 | 12 | 2 | 32 | 15 | 50 | 28 | 28 | 33 | 33 | 8 | 13 | 34 | 35 | 9 | 28 | 29 | 8 | 9 | 13 | 28 | 45 | 7 | 6 | 15 | 26 |
| 215:week_14_arm_1 | 29 | 9 | 8 | 27 | 8 | 8 | 9 | 28 | 25 | 27 | 12 | 0 | 12 | 26 | 17 | 50 | 30 | 30 | 35 | 23 | 10 | 13 | 28 | 37 | 11 | 30 | 31 | 10 | 9 | 15 | 30 | 47 | 9 | 8 | 17 | 28 |
| 215:week_26_arm_1 | 27 | 7 | 6 | 25 | 6 | 6 | 7 | 26 | 23 | 33 | 2 | 12 | 0 | 32 | 15 | 50 | 28 | 28 | 33 | 33 | 8 | 13 | 34 | 35 | 9 | 28 | 29 | 8 | 9 | 13 | 28 | 45 | 7 | 6 | 15 | 26 |
| 215:week_52_arm_1 | 49 | 29 | 28 | 47 | 28 | 28 | 29 | 8 | 13 | 43 | 32 | 26 | 32 | 0 | 21 | 30 | 10 | 10 | 15 | 47 | 30 | 33 | 4 | 57 | 31 | 10 | 55 | 30 | 29 | 35 | 10 | 27 | 29 | 28 | 21 | 8 |
| 215:week_78_arm_1 | 32 | 10 | 11 | 30 | 11 | 11 | 10 | 15 | 10 | 38 | 15 | 17 | 15 | 21 | 0 | 39 | 17 | 17 | 20 | 38 | 13 | 18 | 23 | 40 | 12 | 17 | 42 | 13 | 14 | 16 | 17 | 34 | 10 | 11 | 2 | 15 |
| 257:week_0_arm_1 | 29 | 47 | 46 | 29 | 46 | 46 | 47 | 26 | 31 | 37 | 50 | 50 | 50 | 30 | 39 | 0 | 24 | 24 | 29 | 31 | 44 | 47 | 28 | 37 | 45 | 24 | 37 | 44 | 43 | 49 | 24 | 7 | 47 | 46 | 39 | 26 |
| 257:week_2_arm_1 | 45 | 25 | 24 | 43 | 24 | 24 | 25 | 4 | 9 | 51 | 28 | 30 | 28 | 10 | 17 | 24 | 0 | 2 | 7 | 47 | 22 | 27 | 8 | 49 | 23 | 2 | 51 | 22 | 23 | 27 | 2 | 23 | 25 | 24 | 17 | 4 |
| 257:week_14_arm_1 | 45 | 25 | 24 | 43 | 24 | 24 | 25 | 4 | 9 | 51 | 28 | 30 | 28 | 10 | 17 | 24 | 2 | 0 | 7 | 47 | 22 | 27 | 8 | 49 | 23 | 2 | 51 | 22 | 23 | 27 | 2 | 23 | 25 | 24 | 17 | 4 |
| 257:week_26_arm_1 | 50 | 28 | 29 | 48 | 29 | 29 | 28 | 9 | 12 | 56 | 33 | 35 | 33 | 15 | 20 | 29 | 7 | 7 | 0 | 52 | 27 | 32 | 13 | 46 | 26 | 7 | 56 | 27 | 28 | 22 | 7 | 28 | 28 | 29 | 20 | 9 |
| 275:week_0_arm_1 | 14 | 30 | 29 | 12 | 29 | 29 | 30 | 49 | 46 | 12 | 33 | 23 | 33 | 47 | 38 | 31 | 47 | 47 | 52 | 0 | 27 | 30 | 45 | 18 | 28 | 47 | 12 | 27 | 26 | 32 | 47 | 32 | 30 | 29 | 38 | 49 |
| 275:week_2_arm_1 | 25 | 5 | 4 | 23 | 4 | 4 | 5 | 24 | 21 | 31 | 8 | 10 | 8 | 30 | 13 | 44 | 22 | 22 | 27 | 27 | 0 | 7 | 28 | 29 | 3 | 22 | 31 | 2 | 3 | 7 | 22 | 43 | 5 | 4 | 13 | 24 |
| 275:week_14_arm_1 | 30 | 10 | 9 | 28 | 9 | 9 | 10 | 29 | 26 | 34 | 13 | 13 | 13 | 33 | 18 | 47 | 27 | 27 | 32 | 30 | 7 | 0 | 31 | 34 | 8 | 27 | 34 | 7 | 6 | 12 | 27 | 48 | 10 | 9 | 18 | 29 |
| 275:week_52_arm_1 | 51 | 31 | 30 | 49 | 30 | 30 | 31 | 10 | 15 | 45 | 34 | 28 | 34 | 4 | 23 | 28 | 8 | 8 | 13 | 45 | 28 | 31 | 0 | 55 | 29 | 8 | 53 | 28 | 27 | 33 | 8 | 29 | 31 | 30 | 23 | 10 |
| 305:week_0_arm_1 | 18 | 32 | 31 | 16 | 31 | 31 | 32 | 51 | 48 | 22 | 35 | 37 | 35 | 57 | 40 | 37 | 49 | 49 | 46 | 18 | 29 | 34 | 55 | 0 | 30 | 49 | 22 | 29 | 30 | 26 | 49 | 36 | 32 | 31 | 40 | 51 |
| 305:week_26_arm_1 | 26 | 4 | 5 | 24 | 5 | 5 | 4 | 25 | 20 | 32 | 9 | 11 | 9 | 31 | 12 | 45 | 23 | 23 | 26 | 28 | 3 | 8 | 29 | 30 | 0 | 23 | 32 | 3 | 4 | 6 | 23 | 44 | 4 | 5 | 12 | 25 |
| 305:week_52_arm_1 | 45 | 25 | 24 | 43 | 24 | 24 | 25 | 4 | 9 | 51 | 28 | 30 | 28 | 10 | 17 | 24 | 2 | 2 | 7 | 47 | 22 | 27 | 8 | 49 | 23 | 0 | 51 | 22 | 23 | 27 | 2 | 23 | 25 | 24 | 17 | 4 |
| 365:week_0_arm_1 | 20 | 34 | 33 | 18 | 33 | 33 | 34 | 53 | 50 | 18 | 29 | 31 | 29 | 55 | 42 | 37 | 51 | 51 | 56 | 12 | 31 | 34 | 53 | 22 | 32 | 51 | 0 | 31 | 30 | 36 | 51 | 38 | 34 | 33 | 42 | 53 |
| 365:week_2_arm_1 | 25 | 5 | 4 | 23 | 4 | 4 | 5 | 24 | 21 | 31 | 8 | 10 | 8 | 30 | 13 | 44 | 22 | 22 | 27 | 27 | 2 | 7 | 28 | 29 | 3 | 22 | 31 | 0 | 3 | 7 | 22 | 43 | 5 | 4 | 13 | 24 |
| 365:week_14_arm_1 | 26 | 6 | 5 | 24 | 5 | 5 | 6 | 25 | 22 | 30 | 9 | 9 | 9 | 29 | 14 | 43 | 23 | 23 | 28 | 26 | 3 | 6 | 27 | 30 | 4 | 23 | 30 | 3 | 0 | 8 | 23 | 44 | 6 | 5 | 14 | 25 |
| 365:week_26_arm_1 | 30 | 8 | 9 | 28 | 9 | 9 | 8 | 29 | 24 | 36 | 13 | 15 | 13 | 35 | 16 | 49 | 27 | 27 | 22 | 32 | 7 | 12 | 33 | 26 | 6 | 27 | 36 | 7 | 8 | 0 | 27 | 48 | 8 | 9 | 16 | 29 |
| 365:week_52_arm_1 | 45 | 25 | 24 | 43 | 24 | 24 | 25 | 4 | 9 | 51 | 28 | 30 | 28 | 10 | 17 | 24 | 2 | 2 | 7 | 47 | 22 | 27 | 8 | 49 | 23 | 2 | 51 | 22 | 23 | 27 | 0 | 23 | 25 | 24 | 17 | 4 |
| 527:week_0_arm_1 | 24 | 42 | 41 | 24 | 41 | 41 | 42 | 21 | 26 | 34 | 45 | 47 | 45 | 27 | 34 | 7 | 23 | 23 | 28 | 32 | 43 | 48 | 29 | 36 | 44 | 23 | 38 | 43 | 44 | 48 | 23 | 0 | 42 | 41 | 34 | 21 |
| 527:week_2_arm_1 | 24 | 2 | 3 | 22 | 3 | 3 | 2 | 23 | 18 | 30 | 7 | 9 | 7 | 29 | 10 | 47 | 25 | 25 | 28 | 30 | 5 | 10 | 31 | 32 | 4 | 25 | 34 | 5 | 6 | 8 | 25 | 42 | 0 | 3 | 10 | 23 |
| 527:week_14_arm_1 | 23 | 3 | 2 | 21 | 2 | 2 | 3 | 22 | 19 | 29 | 6 | 8 | 6 | 28 | 11 | 46 | 24 | 24 | 29 | 29 | 4 | 9 | 30 | 31 | 5 | 24 | 33 | 4 | 5 | 9 | 24 | 41 | 3 | 0 | 11 | 22 |
| 527:week_26_arm_1 | 32 | 10 | 11 | 30 | 11 | 11 | 10 | 15 | 10 | 38 | 15 | 17 | 15 | 21 | 2 | 39 | 17 | 17 | 20 | 38 | 13 | 18 | 23 | 40 | 12 | 17 | 42 | 13 | 14 | 16 | 17 | 34 | 10 | 11 | 0 | 15 |
| 527:week_52_arm_1 | 43 | 23 | 22 | 41 | 22 | 22 | 23 | 2 | 7 | 49 | 26 | 28 | 26 | 8 | 15 | 26 | 4 | 4 | 9 | 49 | 24 | 29 | 10 | 51 | 25 | 4 | 53 | 24 | 25 | 29 | 4 | 21 | 23 | 22 | 15 | 0 |
Defining the similarity metric
Use --similarity-metric-cohort to choose the cohort metric. The default value is hamming; the alternative is jaccard.
pheno-ranker -r individuals.json --similarity-metric-cohort jaccard
Sparse Matrix Market output
By default, cohort mode writes a dense tab-separated matrix (matrix.txt). For large cohorts, you can instead write a sparse Matrix Market coordinate file:
pheno-ranker -r individuals.json --matrix-format mtx -o matrix.mtx
The mtx format stores one triangle of the symmetric matrix and writes only non-zero values. It is always RAM-light and does not use the dense in-memory matrix cache controlled by --max-matrix-records-in-ram.
The Matrix Market file includes comment lines mapping 1-based matrix indexes back to individual IDs:
% id 1 107:week_0_arm_1
% id 2 107:week_2_arm_1
Matrix output and Cytoscape graph output are generated independently. This means --matrix-format mtx can be combined with --cytoscape-json.
Exporting intermediate files
It is possible to export all intermediate files, as well as a file indicating coverage, with --export (--e).
Examples:
pheno-ranker -r individuals.json --export
pheno-ranker -r individuals.json --export my_fav_id # choose a prefix
The intermediate files can be used for further processing (e.g., import to a database; see FAQs) or to make informed decisions. For instance, the file export.coverage_stats.json has stats on the coverage of each term (1D-key) in the cohort. It is possible to go more granular with a tool like jq that parses JSON. For instance:
jq -r 'to_entries | map(.key + ": " + (.value | length | tostring))[]' < export.ref_hash.json
This command will print how many variables per individual were actually used to perform the comparison. You can post-process the output to check for unbalanced data.
Included R scripts
You can find in the link below a few examples to perform clustering and multidimensional scaling with your data:
Clustering
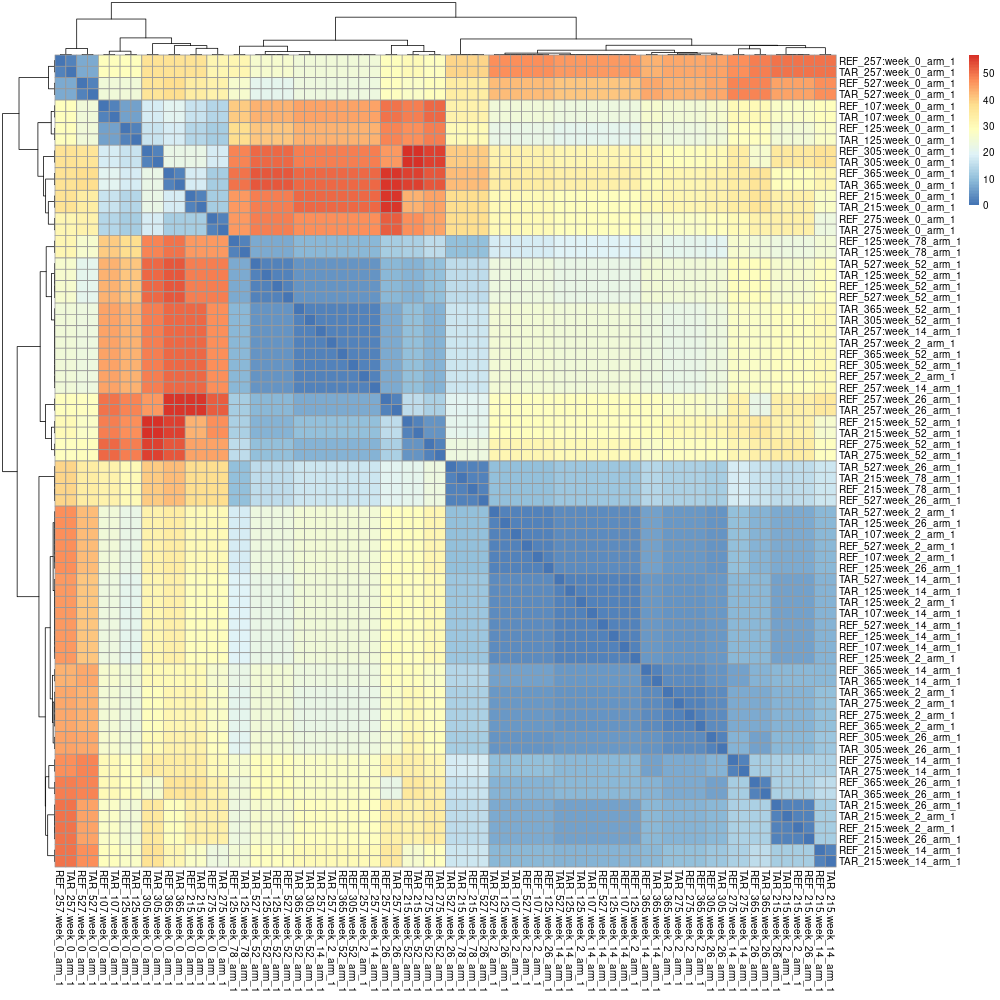
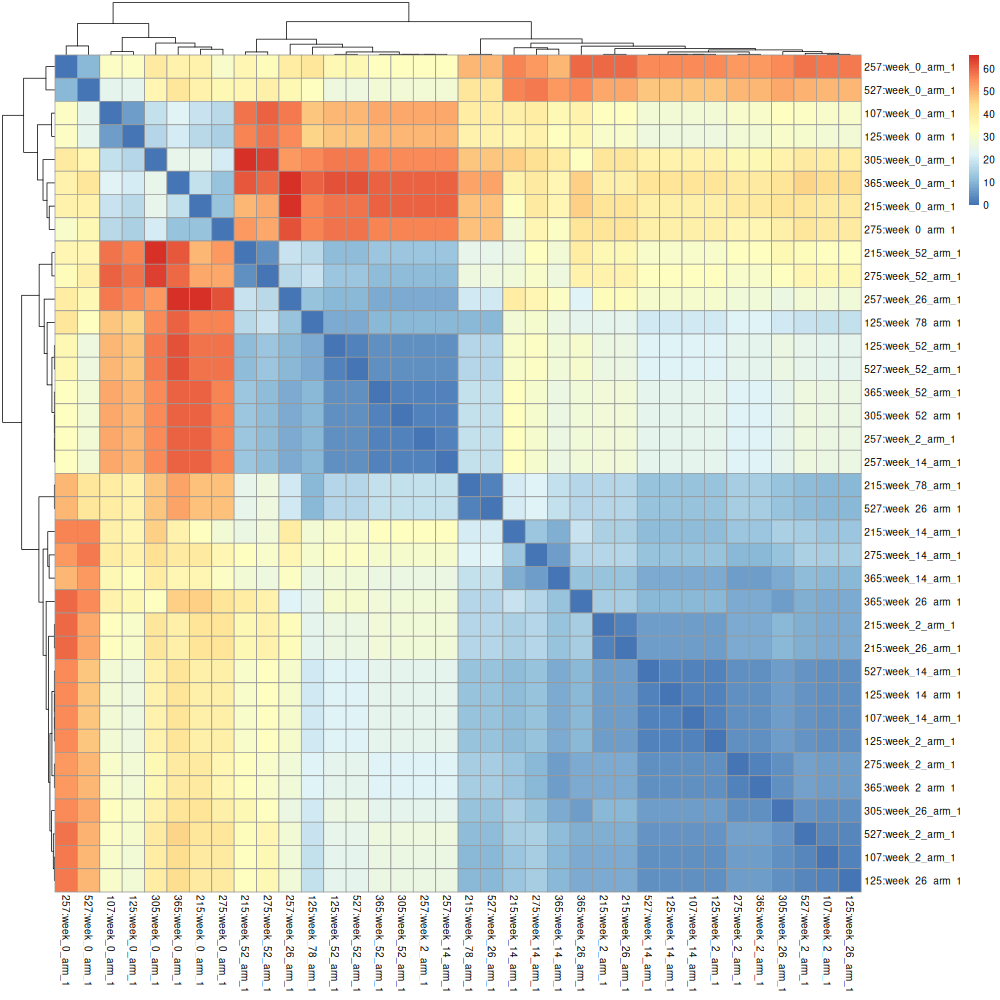
The matrix can be processed to obtain a heatmap:
R code
# Load library
library("pheatmap")
#library("heatmaply") # could not install
# Read in the input file as a matrix
data <- as.matrix(read.table("matrix.txt", header = TRUE, row.names = 1, check.names = FALSE))
# Save image
png(filename = "heatmap.png", width = 1000, height = 1000,
units = "px", pointsize = 12, bg = "white", res = NA)
# Create the heatmap with row and column labels
#heatmap(data, Rowv = FALSE, Colv = FALSE, labRow = rownames(data), labCol = colnames(data))
pheatmap(data)
#heatmaply(data)
#dev.off()
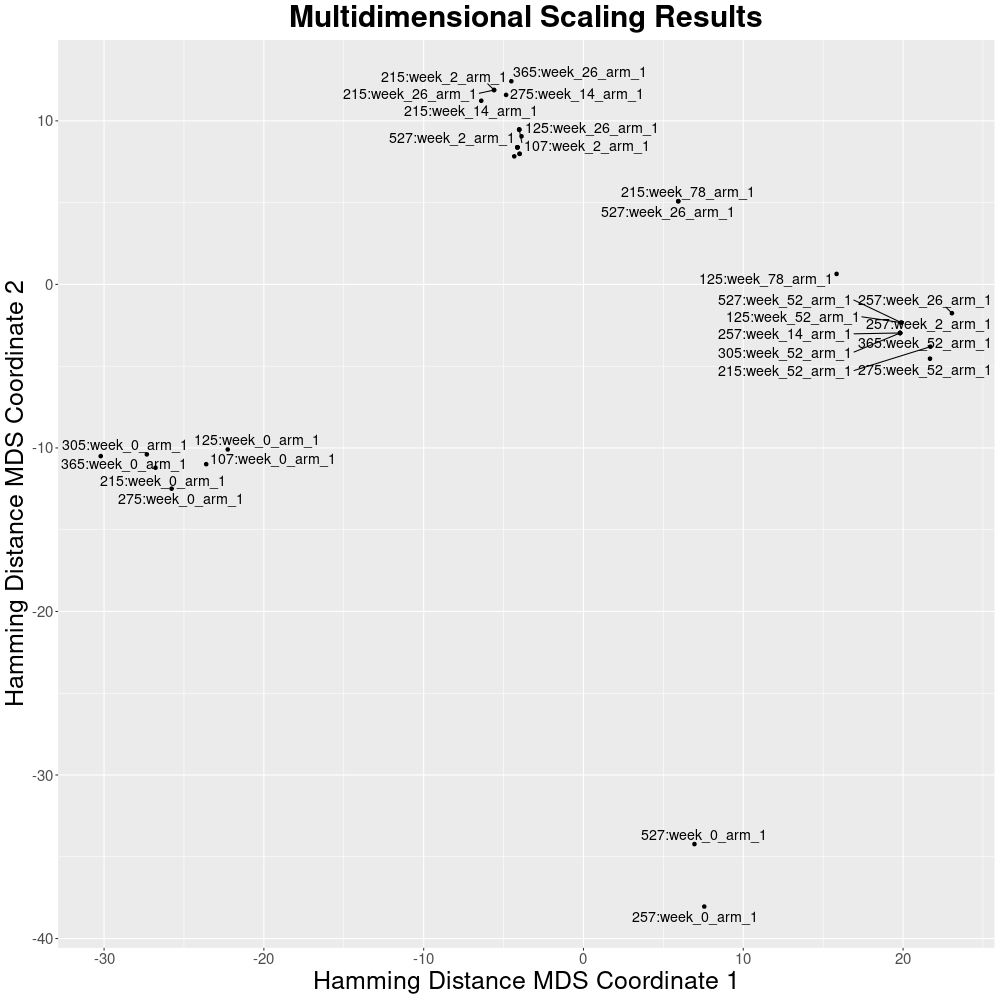
Dimensionality reduction
The same matrix can be processed with multidimensional scaling to reduce the dimensionality.
R code
library(ggplot2)
library(ggrepel)
# Read in the input file as a matrix
data <- as.matrix(read.table("matrix.txt", header = TRUE, row.names = 1, check.names = FALSE))
# Calculate distance matrix
#d <- dist(data)
#d <- 1 - data # J-similarity to J-distance
# Perform multidimensional scaling
#fit <- cmdscale(d, eig=TRUE, k=2)
fit <- cmdscale(data, eig=TRUE, k=2)
# Extract (x, y) coordinates of multidimensional scaling
x <- fit$points[,1]
y <- fit$points[,2]
# Create data frame
df <- data.frame(x, y, label=row.names(data))
# Save image
png(filename = "mds.png", width = 1000, height = 1000,
units = "px", pointsize = 12, bg = "white", res = NA)
# Create scatter plot
ggplot(df, aes(x, y, label = label)) +
geom_point() +
geom_text_repel(size = 5, # Adjust the size of the text
box.padding = 0.2, # Adjust the padding around the text
max.overlaps = 10) + # Change the maximum number of overlaps
labs(title = "Multidimensional Scaling Results",
x = "Hamming Distance MDS Coordinate 1",
y = "Hamming Distance MDS Coordinate 2") + # Add title and axis labels
theme(
plot.title = element_text(size = 30, face = "bold", hjust = 0.5),
axis.title = element_text(size = 25),
axis.text = element_text(size = 15))
#dev.off()
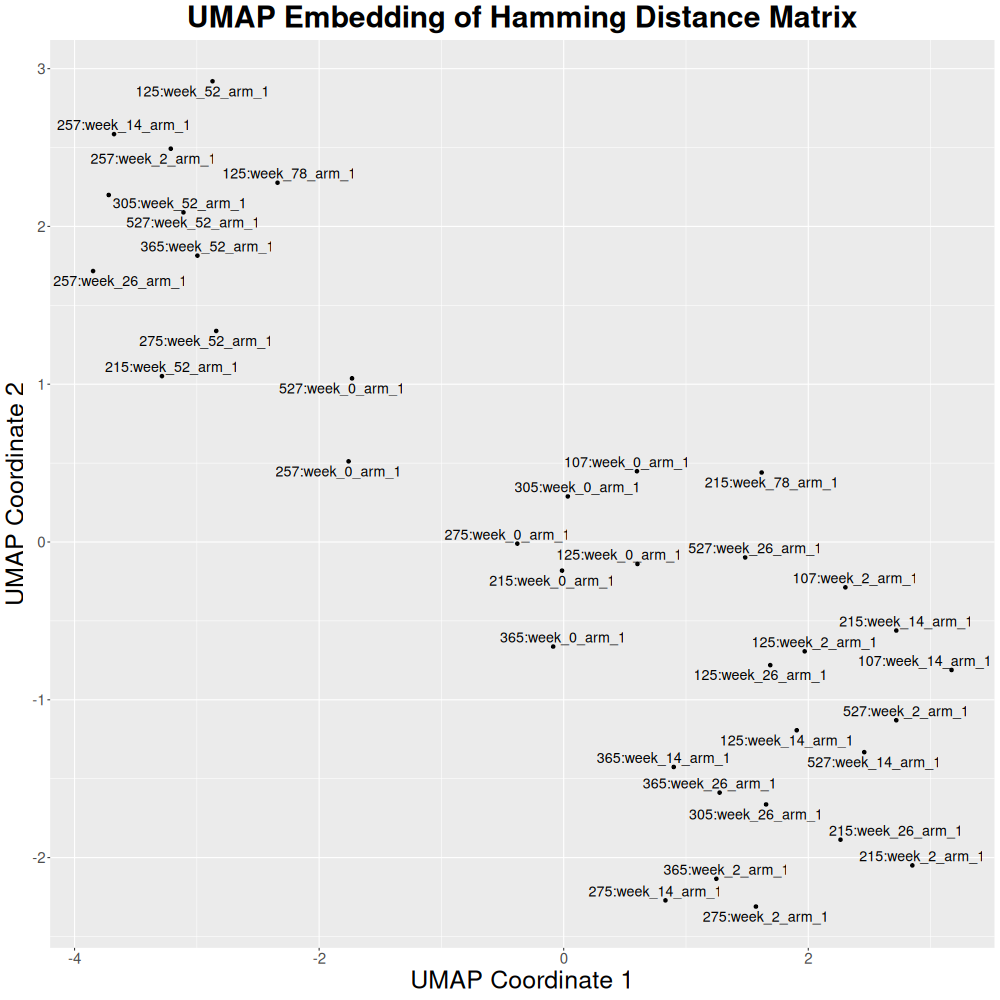
Or the dimensionality can be reduced with UMAP:
R code
# -- Install uwot on the fly if needed
if (!requireNamespace("uwot", quietly = TRUE)) {
install.packages("uwot", repos="https://cloud.r-project.org")
}
# -- Load libraries
library(uwot)
library(ggplot2)
library(ggrepel)
# -- Read in the input file as a full distance matrix
data <- as.matrix(
read.table("matrix.txt",
header=TRUE,
row.names=1,
check.names=FALSE)
)
# -- Convert to a 'dist' object so uwot knows these are distances
d <- as.dist(data)
# -- Set seed for reproducibility
set.seed(42)
# -- Run UMAP directly on the distances
# Passing a 'dist' object lets uwot build the k-NN graph from your distances
umap_res <- umap(
d,
n_neighbors=30,
min_dist=0.3,
n_components=2
)
# -- Extract UMAP coordinates
x <- umap_res[,1]
y <- umap_res[,2]
# -- Build a data frame for plotting
df <- data.frame(
x=x,
y=y,
label=rownames(data)
)
# -- Open PNG device
png(filename="umap.png",
width=1000,
height=1000,
units="px",
pointsize=12,
bg="white",
res=NA)
# -- Create scatter plot with labels
ggplot(df, aes(x=x, y=y, label=label)) +
geom_point() +
geom_text_repel(
size=5,
box.padding=0.2,
max.overlaps=10
) +
labs(
title="UMAP Embedding of Hamming Distance Matrix",
x="UMAP Coordinate 1",
y="UMAP Coordinate 2"
) +
theme(
plot.title=element_text(size=30, face="bold", hjust=0.5),
axis.title=element_text(size=25),
axis.text=element_text(size=15)
)
# -- Close the device
dev.off()
Graph analytics
Pheno-Ranker has an option for creating a graph in JSON format, compatible with the Cytoscape ecosystem.
Bash code for Cytoscape-compatible graph/network
pheno-ranker -r individuals.json --cytoscape-json
This command generates a graph.json file, as well as a matrix.txt file. The graph is generated directly from the binary comparison hashes, not by parsing the matrix file, so it can also be combined with Matrix Market output:
pheno-ranker -r individuals.json --matrix-format mtx -o matrix.mtx --cytoscape-json graph.json
Large graphs can be filtered by edge weight:
# Hamming distance: keep close pairs
pheno-ranker -r individuals.json --cytoscape-json --graph-max-weight 10
# Jaccard similarity: keep highly similar pairs
pheno-ranker -r individuals.json --similarity-metric-cohort jaccard --cytoscape-json --graph-min-weight 0.7
To produce summary statistics, use:
pheno-ranker -r individuals.json --cytoscape-json --graph-stats
This command will produce a file called graph_stats.txt. For additional information, see the generic JSON tutorial.
We use individuals.json again, but pass it twice to simulate two cohorts. Pheno-Ranker adds a CX_ prefix to the primary_key values so each record can be traced back to its source cohort.
pheno-ranker -r individuals.json individuals.json
Absolutely, a cohort can indeed be composed of a single individual. This allows for an analysis involving both a cohort and specific patient(s) simultaneously.
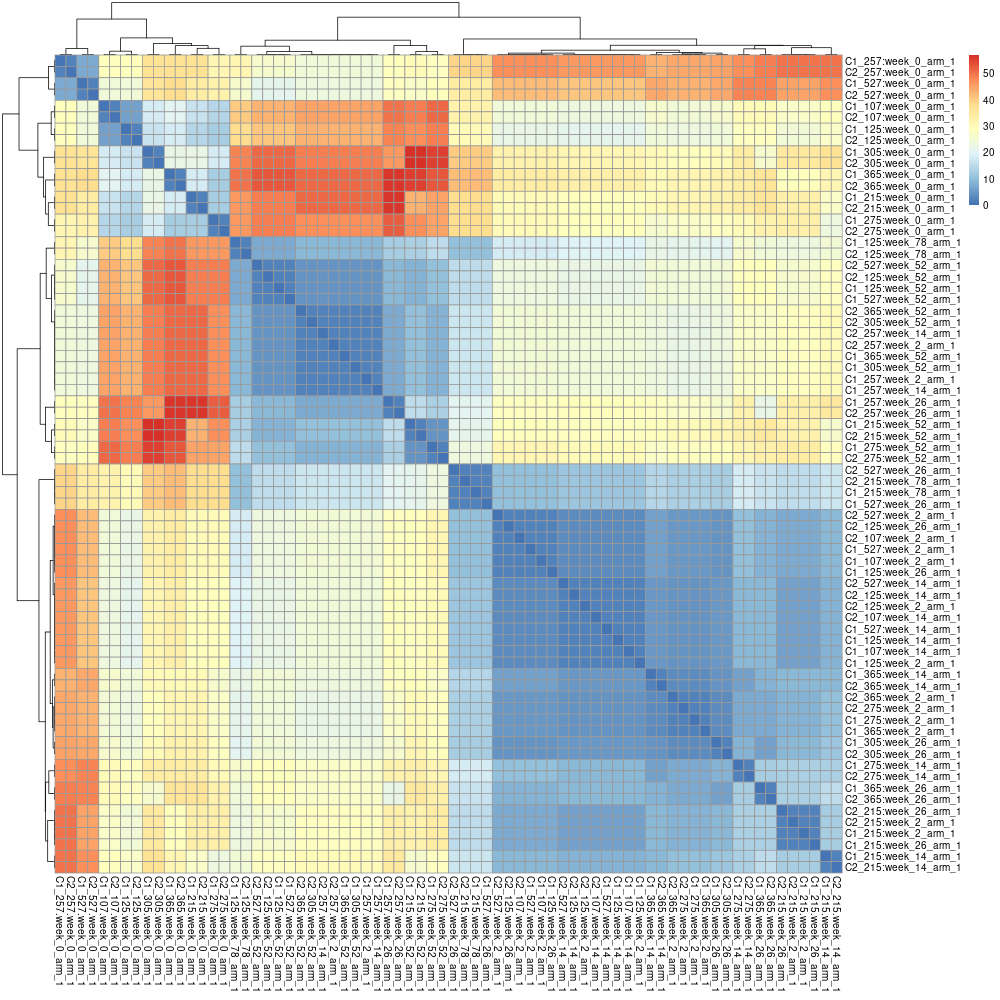
The prefixes can be changed with the flag --append-prefixes:
pheno-ranker -r individuals.json individuals.json --append-prefixes REF TAR
This will create a matrix.txt file of (36+36) x (36+36) cells. Again, this matrix can be processed with R: